This one is for the science teachers, and all other teachers too!
Thursday, July 31, 2014
Friday, July 25, 2014
Small Things, Big Problem: Microplastics Uptake in Shore Crabs
Lately I've been gearing up for some nano-particle research, and so I've been doing a lot of reading about very small things. While perusing the literature, I came across a paper published online in Environmental Science and Technology that takes a look at microplastics.
Let’s start with the Great Pacific Garbage Patch, a very good example of this type of marine pollution. This huge collection of marine debris in the North Pacific Ocean is created by an ocean gyre, a stable circular ocean current that draws in debris where it is trapped and builds up. The collected debris is our litter – plastics and other material that are not biodegradable. They can’t escape the gyre, they just collect. And as they sit out there swirling around, they break down into smaller and smaller pieces called microplastics.
Microplastics are defined as those plastic particles less than 5 mm in length, and these small particles are a huge marine pollution problem. They are classified into two groups: (1) primary microplastics that are created at the microscale for use in products like cosmetics and drugs and (2) secondary microplastics that are products of the breakdown of larger items. As a whole, they are persistent and widespread – we’re talking worldwide, the Great Pacific Garbage Patch is just the most well-known aggregation. These microplastics are very abundant, we’re talking 1,000-100,000 particles per cubic meter of seawater! And there is growing evidence of the danger these tiny materials are having on marine life, everything from turtles to sea birds to fish and even zooplankton.
A new study by Watts et al. takes look at the uptake of these microplastics in the shore crab (Carcinus maenas). Previous studies have shown that an important prey species of the shore crab, the common mussel (Mytilus edulis), accumulates microplastics as it filters the water for food (“ventilation”). In laboratory conditions, the direct transfer of microplastics from mussels to crabs has been shown, but then again, it has also been shown that crabs uptake microplastics as they pull water through their gills. So what exactly is going on here? How are these crabs exposed and are they able to clear the microplastics from their bodies?
This is one of those studies where I just love to describe the methods. The first thing the researchers had to do was to assess the ability of the crabs to uptake microplastics (in the form of 8-10 um polystyrene microspheres) through their gills. To do this, they fitted the crabs with masks designed to allow measurements of ventilation. Yep, they put little masks on crabs. Picture that. I love science. Next they assessed the ability of the crab to take up microspheres in their food by exposing mussels and then turning them into “jellified mussel homogenate” to then feed to the crabs. I wonder which undergrad had the lovely job of making gelatin mussel popsicles? To see if the microplastcs were cleared, they let the crabs sit in their tanks and tested the abundance of microspheres in the water during water changes every 2 days for 22 days, sampling periodically. During each stage of the experiment, they measured the abundance of microspheres in the gut and gill tissues, fecal material, and hemolymph (like blood). Using fluorescent microscopy and Coherent Raman scattering microscopy (CRS; a multiphoton microscopy that produces label-free contrast of both the target sample and the surrounding biological matrix), they were able to look at the location of the microplastics within the tissues.
The researchers found that the masked crabs took up 31,000-62,000 microspheres (0.39-7.7% of the initial exposure concentration) into their gills after only 16 hours. But this uptake was not even across the gills, with greater uptake in the posterior gills. The crabs where able to expel some of the spheres, but slowly, still expiring microspheres 21 days after being exposed. Imaging the gills showed the microspheres to be associated with the gill epidermis. The feeding experiment showed all crabs to have microspheres in their foregut and later in their fecal material. The residence time of these microspheres was short, but still took longer to excrete than regular food waste, up to 14 days. Microscopy showed microspheres associated with the internal setae of the foregut lining. But, neither the ventilation experiment nor the feeding experiment showed any microspheres in the hemolymph.
Back to the question of what’s going on here? The shore crabs did take up microplastics in both types of exposure, but residence time is the key. They were able to clear the microplastics they got through dietary means, but they were still trying to clear microplastics they took up during ventilation almost a month later. The authors constructed a model to explain the mechanism of the movement of the microplastics. They found that the crabs tended to exhibit an asymmetry in microplastic uptake in the gills, which they attributed to the pumping mechanism of the scaphognathite being more dominant on one side of the gill chamber. Also, the posterior gills have a larger surface area than the anterior gills so they are more likely to take up microplastics into their lamellae. The crabs were unable to dislodge the tiny particles by normal gill cleaning actions. It is interesting that no microspheres were found in the hemolymph at any of the sampling points in either experiment. That suggests that there is no movement of the particles. It is likely that the particle size they used (10 um) was a little bit too large as it has been shown that sizes of 0.5 um are able to translocate in these crabs. This idea of particle size is something I’ve been seeing with increasing frequency within the nano-particle literature, along with polymer type, shape, and coatings. To that I would add that species is probably also in the mix as gills in crabs and fish are structured differently, and nano-particles have been shown to move in to organs like the liver in fish.
Studies like this are interesting because they show how very small things can become a very large problem affecting multiple tissues of the same organism up to multiple levels of a trophic cascade. I mean, think about it, even we humans could be affected. After all, we consume a lot of crab. How many microplastics are you ingesting when you stop at the crab shack for a quick lunch?
I know I used some technical terms, if you need some help with crustacean anatomy check out Invertebrate Anatomy OnLine.
(image via UGA Evolution 3000H)
Labels:
arthropods,
crustaceans,
invertebrates,
oceanography
Thursday, July 24, 2014
Wednesday, July 23, 2014
Let it Go, Med School
Apparently med school students just love to film parodies. A lot.
Here are a couple of good ones using songs from Frozen.
Here are a couple of good ones using songs from Frozen.
Thursday, July 17, 2014
Origami Science
Origami...one sheet, no cuts. Tough. Now, what does science have to do with origami? As it turns out, a lot.
Robert J. Lang is a former NASA laser physicist who combines his knowledge of math and science to create origami masterpieces. His pieces are known for their great detail and realism, and they are some of the most complex origami designs ever created.
Lang groups the origami science into three categories:
Robert J. Lang is a former NASA laser physicist who combines his knowledge of math and science to create origami masterpieces. His pieces are known for their great detail and realism, and they are some of the most complex origami designs ever created.
Lang groups the origami science into three categories:
- Origami Mathematics - This is a subset of mathematics using one of the simplest sets of operations called "Huzita-Justin Axioms." This os set of six (possibly seven?) distinctly different ways to create a single crease by alligning one or more combinations of points and lines expanded with origami geometric constructions that solve for an exact angle quintisection.
- Computational Origami - This is a subset of computer science that uses computational geometry, and the associated algorithms, as a tool for origami design and computations.
- Origami Technology - This is the application of origami techniques to real-world solution making problems in fields like engineering and industrial design. Think: How to fold an car's airbag or folding a telescope.
Using these techniques, Land has been able to create more than 500 original origami compositions using different types of paper and other materials like bronze and self-folding polymers. He has written 14 books on origami and designed origami software offered for free on his website. Lang has sown at some pretty big venues too, including the and the Shumei Hall Gallery in Pasadena, California, Cooper Union in New York, the Santa Fe Botanical Garden in Santa Fe, New Mexico, New York's Fashion Institute of Technology, etc. etc. Currently, over 100 of his pieces are on display at a solo show, Folded, at the Williamson Gallery at the Art Center College of Design, in Pasadena, California.
Here are some of my favorites:.
Stories that include more photographs and videos of Lang's wonderful origami creations:
Robert J. Lang's Origami Website
Fast Company "Art Folds into Science in Robert Lang's Extreme Origami"
Williamson Gallery's Information on "Folded: The Origami of Robert J. Lang"
Science Alert "Behind the science of Robert J. Lang's origami"
Here are some of my favorites:.
 |
| PRAYING MANTIS, OPUS 416 Composed and folded: 2012; One uncut square of paper 4" Image: Emre Ayaroglu/Flickr via Science Alert |
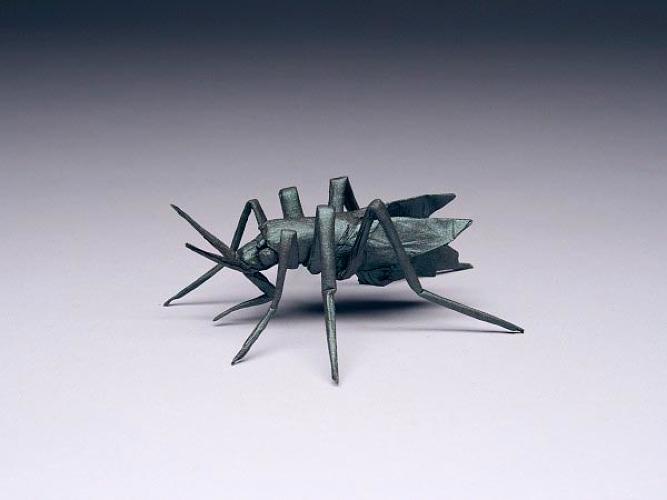 |
| AEDES AGEYPTI, OPUS 619 Composed and folded: 2012, Commissioned by The New Yorker Magazine c/o Robert Lang via Fast Company |
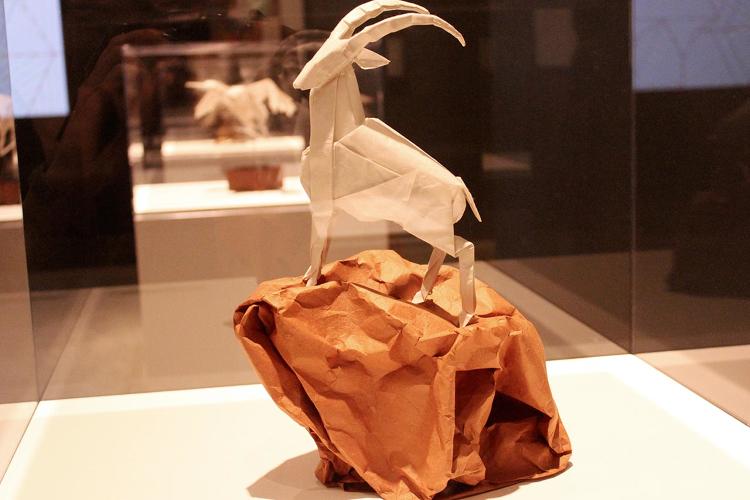 |
| THE SENTINEL II, OPUS 627 Composed and folded: 2012; Two uncut squares of Korean hanji, 14" Photo by Susan Karlin via Fast Company |
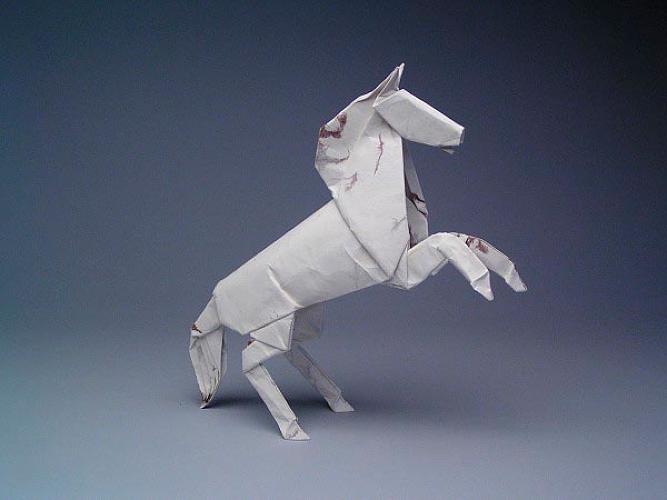 |
| ALAMO STALLION, OPUS 384 Composed: 2001, folded 2012; One uncut square of Origamido paper 12" c/o Robert Lang via Fast Company |
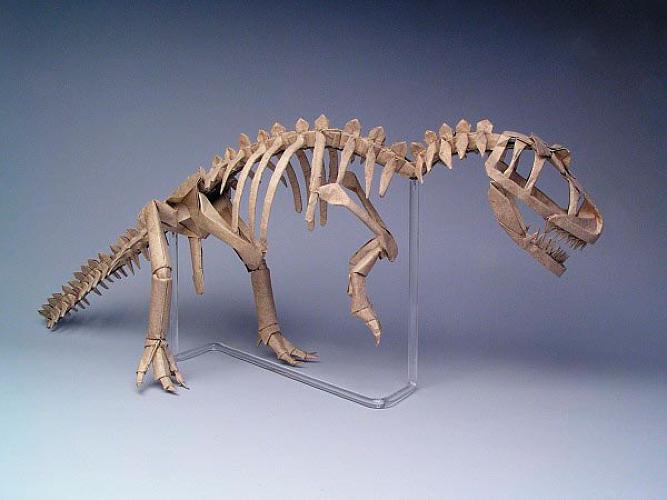 |
| ALLOSAURUS SKELETON, OPUS 326 16 uncut squares of Wyndstone Marble paper 24" c/o Robert Lang via Fast Company |
Stories that include more photographs and videos of Lang's wonderful origami creations:
Robert J. Lang's Origami Website
Fast Company "Art Folds into Science in Robert Lang's Extreme Origami"
Williamson Gallery's Information on "Folded: The Origami of Robert J. Lang"
Science Alert "Behind the science of Robert J. Lang's origami"
Tuesday, July 15, 2014
Breaking Up is Hard to Do: Photosynthesis, Water-Splitting, and the OEC
A very very cool paper was recently published online. The paper details a study that shows the first images of water splitting apart during photosynthesis. So pick you jaw up off the table and we’ll get into the nitty-gritty details.
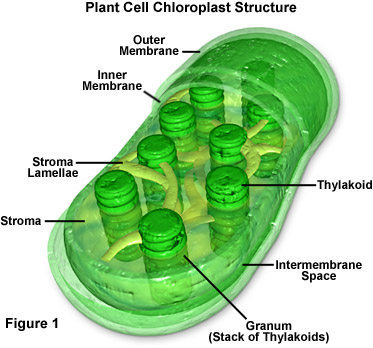 Let’s start by accessing your long-term memory, dragging out some of that basic biology information you buried after high school and grabbing on to that dusty file about photosynthesis. If you remember, plants have little green, bean-shaped energy factories in their cells called chloroplasts. These chloroplasts are filled thylakoids stacked up in grana. The thylakoid membranes contain networks of pigments, including chlorophyll, arranged in aggregates or complexes. Think of them kinda like light energy harvesters. Energy is captured for functional and structural units of protein complexes called Photosystem I (PSI) and Photosystem II (PSII). PSI is the light reaction and converts light energy to chemical energy. The pigments of the complexes each absorb light and then pass along that light energy to the central chlorophyll molecule to do photosynthesis. The energy obtained in this reaction is stored in ATP (adenosine triphosphate) and NADPH (nicotinamide adenine dinucleotide phosphate-oxidase) molecules. PSII, the dark reaction, takes place in the stroma within the chloroplast. This reaction uses the Calvin cycle to convert carbon dioxide and energy from ATP into glucose (sugar). To say that is photosynthesis put shortly and simply would be an understatement, but keep this basic reaction in mind:
Let’s start by accessing your long-term memory, dragging out some of that basic biology information you buried after high school and grabbing on to that dusty file about photosynthesis. If you remember, plants have little green, bean-shaped energy factories in their cells called chloroplasts. These chloroplasts are filled thylakoids stacked up in grana. The thylakoid membranes contain networks of pigments, including chlorophyll, arranged in aggregates or complexes. Think of them kinda like light energy harvesters. Energy is captured for functional and structural units of protein complexes called Photosystem I (PSI) and Photosystem II (PSII). PSI is the light reaction and converts light energy to chemical energy. The pigments of the complexes each absorb light and then pass along that light energy to the central chlorophyll molecule to do photosynthesis. The energy obtained in this reaction is stored in ATP (adenosine triphosphate) and NADPH (nicotinamide adenine dinucleotide phosphate-oxidase) molecules. PSII, the dark reaction, takes place in the stroma within the chloroplast. This reaction uses the Calvin cycle to convert carbon dioxide and energy from ATP into glucose (sugar). To say that is photosynthesis put shortly and simply would be an understatement, but keep this basic reaction in mind:
6 CO2 + 6 H2O à C6H12O6 + 6 O2
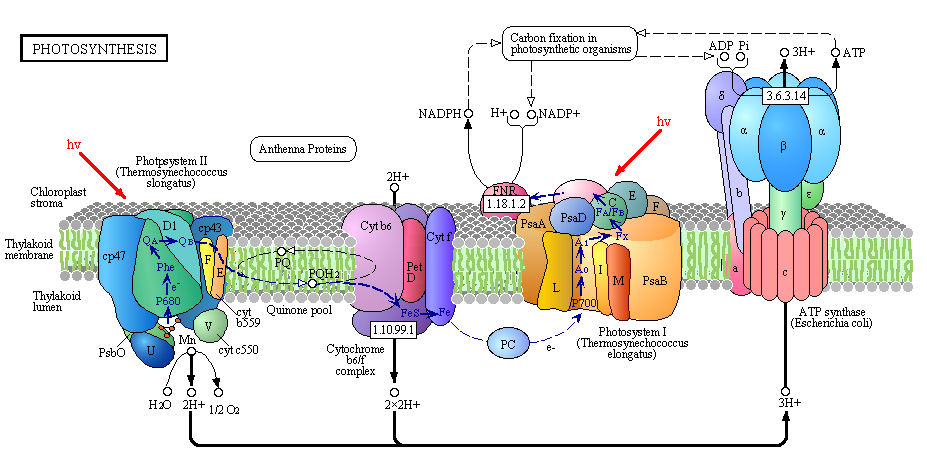
It is important to mention that in PSII, water is photochemically oxidized to dioxygen (O2) by the oxygen-evolving complex (OEC), a metalloenzyme cluster containing manganese and calcium. The OEC cycles through five photo-catalytic stages (S0-S4) in which four electrons are sequentially extracted from the OEC in four light-driven charge-separation events by a repeatedly photo-oxidized chlorophyll center (Kok cycle). This is the reaction that makes all that oxygen we breathe.
2 H2O à S0-S4 à O2 + 4 H+ + 4 e-
The new paper by Kupitz et al. (and al. and al. and al.) published in Nature looks closer (very close!) at this PSII water-splitting reaction. They had some issues to overcome if they wanted to collect more information on this reaction, mostly involving the static nature of X-ray crystallography and the damage done to the OEC with this method. Traditional X-ray crystallography enables 1.9Å resolution (near atomic) but the OEC probably suffers X-ray damage. To overcome this, the researchers used serial femtosecond crystallography. This method uses single shot diffraction patterns are collected from a stream of nanocrystals, using 120 Hz femtosecond (one millionth of a nanosecond!!) pulses from an X-ray Free Electron Laser (XFEL). The second is the quality of the structural information. These pulses are so intense that the sample/specimen is destroyed, but the pulse duration is so short that the diffraction is observed before the destruction occurs. The method produces millions of “snapshots” in hours and can collect time-resolved data for dynamic processes like water oxidation in PSII.
The researchers developed a multiple-laser illumination scheme to observe this dynamic reaction in thermophilic cyanobacterium (Thermosynechococcus elongates). They progressively excited the OEC in dark-adapted PSII nano/microcrystals by two laser pulses from the dark S1 state via the S2 state to the double-flash putative S3 state (5 and 5.5Å resolution). Believe it or not, that was their method put simply. Essentially, they were able to determine the structures of the states and to produce maps of the protein subunits and cofactors of PSII, including the electron transport chain. They found that PSII undergoes significant conformational changes electron acceptor side and at the Mn4CaO5 core of the OEC. The metal cluster significantly elongates, making room and allowing for binding of the incoming water molecules. Then voilà! Water splitting!
So I know what you may be thinking: Why all of that lead-up to a simple protein shape change conclusion? Well, it’s all about mechanism, figuring out the process of photosynthesis at its most basic level. If you think about it, photosynthesis is the biological reaction. It is fundamental to life on Earth as we know it. It converted the oxygen-poor atmosphere of early Earth to the oxygen-rich atmosphere we (and all other respiring organisms) depend on, and continues to supply us with life-giving oxygen. That oxygen comes from this water splitting reaction, and the OEC is one of those structures where you usually have to but "possible model of..." in front. This type of study gives incredible resolution of this structure as well as a new methodology to gain further knowledge. With a more technological viewpoint, work like this could eventually lead to the development of an artificial leaf and synthetic photosynthesis. And, let’s face it, that is really really cool.
 Kupitz, C., Basu, S., Grotjohann, I., Fromme, R., Zatsepin, N., Rendek, K., Hunter, M., Shoeman, R., White, T., Wang, D., James, D., Yang, J., Cobb, D., Reeder, B., Sierra, R., Liu, H., Barty, A., Aquila, A., Deponte, D., Kirian, R., Bari, S., Bergkamp, J., Beyerlein, K., Bogan, M., Caleman, C., Chao, T., Conrad, C., Davis, K., Fleckenstein, H., Galli, L., Hau-Riege, S., Kassemeyer, S., Laksmono, H., Liang, M., Lomb, L., Marchesini, S., Martin, A., Messerschmidt, M., Milathianaki, D., Nass, K., Ros, A., Roy-Chowdhury, S., Schmidt, K., Seibert, M., Steinbrener, J., Stellato, F., Yan, L., Yoon, C., Moore, T., Moore, A., Pushkar, Y., Williams, G., Boutet, S., Doak, R., Weierstall, U., Frank, M., Chapman, H., Spence, J., & Fromme, P. (2014). Serial time-resolved crystallography of photosystem II using a femtosecond X-ray laser Nature DOI: 10.1038/nature13453
Kupitz, C., Basu, S., Grotjohann, I., Fromme, R., Zatsepin, N., Rendek, K., Hunter, M., Shoeman, R., White, T., Wang, D., James, D., Yang, J., Cobb, D., Reeder, B., Sierra, R., Liu, H., Barty, A., Aquila, A., Deponte, D., Kirian, R., Bari, S., Bergkamp, J., Beyerlein, K., Bogan, M., Caleman, C., Chao, T., Conrad, C., Davis, K., Fleckenstein, H., Galli, L., Hau-Riege, S., Kassemeyer, S., Laksmono, H., Liang, M., Lomb, L., Marchesini, S., Martin, A., Messerschmidt, M., Milathianaki, D., Nass, K., Ros, A., Roy-Chowdhury, S., Schmidt, K., Seibert, M., Steinbrener, J., Stellato, F., Yan, L., Yoon, C., Moore, T., Moore, A., Pushkar, Y., Williams, G., Boutet, S., Doak, R., Weierstall, U., Frank, M., Chapman, H., Spence, J., & Fromme, P. (2014). Serial time-resolved crystallography of photosystem II using a femtosecond X-ray laser Nature DOI: 10.1038/nature13453
Arizona State University Science and Tech press release: "ASU-led study yields first snapshots of water splitting in photosynthesis"
Science Daily article: “First snapshots of water splitting in photosynthesis”
For more on the plant physiology:
ASU's page "What is Photosynthesis?"
James Johnson @ FSU page on "The Manganese-calcium oxide cluster of Photosystem II"
Dr. Jakubowski at Saint Johns University "Chapter 8 - Oxidation/Phosphorylation"
(images via FSU Molecular Expressions, Jakubowski's website, and Arizona State University, respectively)
 Let’s start by accessing your long-term memory, dragging out some of that basic biology information you buried after high school and grabbing on to that dusty file about photosynthesis. If you remember, plants have little green, bean-shaped energy factories in their cells called chloroplasts. These chloroplasts are filled thylakoids stacked up in grana. The thylakoid membranes contain networks of pigments, including chlorophyll, arranged in aggregates or complexes. Think of them kinda like light energy harvesters. Energy is captured for functional and structural units of protein complexes called Photosystem I (PSI) and Photosystem II (PSII). PSI is the light reaction and converts light energy to chemical energy. The pigments of the complexes each absorb light and then pass along that light energy to the central chlorophyll molecule to do photosynthesis. The energy obtained in this reaction is stored in ATP (adenosine triphosphate) and NADPH (nicotinamide adenine dinucleotide phosphate-oxidase) molecules. PSII, the dark reaction, takes place in the stroma within the chloroplast. This reaction uses the Calvin cycle to convert carbon dioxide and energy from ATP into glucose (sugar). To say that is photosynthesis put shortly and simply would be an understatement, but keep this basic reaction in mind:
Let’s start by accessing your long-term memory, dragging out some of that basic biology information you buried after high school and grabbing on to that dusty file about photosynthesis. If you remember, plants have little green, bean-shaped energy factories in their cells called chloroplasts. These chloroplasts are filled thylakoids stacked up in grana. The thylakoid membranes contain networks of pigments, including chlorophyll, arranged in aggregates or complexes. Think of them kinda like light energy harvesters. Energy is captured for functional and structural units of protein complexes called Photosystem I (PSI) and Photosystem II (PSII). PSI is the light reaction and converts light energy to chemical energy. The pigments of the complexes each absorb light and then pass along that light energy to the central chlorophyll molecule to do photosynthesis. The energy obtained in this reaction is stored in ATP (adenosine triphosphate) and NADPH (nicotinamide adenine dinucleotide phosphate-oxidase) molecules. PSII, the dark reaction, takes place in the stroma within the chloroplast. This reaction uses the Calvin cycle to convert carbon dioxide and energy from ATP into glucose (sugar). To say that is photosynthesis put shortly and simply would be an understatement, but keep this basic reaction in mind:6 CO2 + 6 H2O à C6H12O6 + 6 O2

It is important to mention that in PSII, water is photochemically oxidized to dioxygen (O2) by the oxygen-evolving complex (OEC), a metalloenzyme cluster containing manganese and calcium. The OEC cycles through five photo-catalytic stages (S0-S4) in which four electrons are sequentially extracted from the OEC in four light-driven charge-separation events by a repeatedly photo-oxidized chlorophyll center (Kok cycle). This is the reaction that makes all that oxygen we breathe.
2 H2O à S0-S4 à O2 + 4 H+ + 4 e-
 |
| Photo by: Mary Zhu @ ASU |
The new paper by Kupitz et al. (and al. and al. and al.) published in Nature looks closer (very close!) at this PSII water-splitting reaction. They had some issues to overcome if they wanted to collect more information on this reaction, mostly involving the static nature of X-ray crystallography and the damage done to the OEC with this method. Traditional X-ray crystallography enables 1.9Å resolution (near atomic) but the OEC probably suffers X-ray damage. To overcome this, the researchers used serial femtosecond crystallography. This method uses single shot diffraction patterns are collected from a stream of nanocrystals, using 120 Hz femtosecond (one millionth of a nanosecond!!) pulses from an X-ray Free Electron Laser (XFEL). The second is the quality of the structural information. These pulses are so intense that the sample/specimen is destroyed, but the pulse duration is so short that the diffraction is observed before the destruction occurs. The method produces millions of “snapshots” in hours and can collect time-resolved data for dynamic processes like water oxidation in PSII.
The researchers developed a multiple-laser illumination scheme to observe this dynamic reaction in thermophilic cyanobacterium (Thermosynechococcus elongates). They progressively excited the OEC in dark-adapted PSII nano/microcrystals by two laser pulses from the dark S1 state via the S2 state to the double-flash putative S3 state (5 and 5.5Å resolution). Believe it or not, that was their method put simply. Essentially, they were able to determine the structures of the states and to produce maps of the protein subunits and cofactors of PSII, including the electron transport chain. They found that PSII undergoes significant conformational changes electron acceptor side and at the Mn4CaO5 core of the OEC. The metal cluster significantly elongates, making room and allowing for binding of the incoming water molecules. Then voilà! Water splitting!
So I know what you may be thinking: Why all of that lead-up to a simple protein shape change conclusion? Well, it’s all about mechanism, figuring out the process of photosynthesis at its most basic level. If you think about it, photosynthesis is the biological reaction. It is fundamental to life on Earth as we know it. It converted the oxygen-poor atmosphere of early Earth to the oxygen-rich atmosphere we (and all other respiring organisms) depend on, and continues to supply us with life-giving oxygen. That oxygen comes from this water splitting reaction, and the OEC is one of those structures where you usually have to but "possible model of..." in front. This type of study gives incredible resolution of this structure as well as a new methodology to gain further knowledge. With a more technological viewpoint, work like this could eventually lead to the development of an artificial leaf and synthetic photosynthesis. And, let’s face it, that is really really cool.
Arizona State University Science and Tech press release: "ASU-led study yields first snapshots of water splitting in photosynthesis"
Science Daily article: “First snapshots of water splitting in photosynthesis”
For more on the plant physiology:
ASU's page "What is Photosynthesis?"
James Johnson @ FSU page on "The Manganese-calcium oxide cluster of Photosystem II"
Dr. Jakubowski at Saint Johns University "Chapter 8 - Oxidation/Phosphorylation"
(images via FSU Molecular Expressions, Jakubowski's website, and Arizona State University, respectively)
Friday, July 11, 2014
Talk Nerdy to Me
Happy Friday! Here's great Jason Derulo parody for you.
Tuesday, July 8, 2014
You Know You've Worked Too Long in a Lab When
Monday, July 7, 2014
Planetary Beer
Mmmmm.....beer.
Over the past few years I been cultivating my beer snobbery. Long gone are the days when I would drink for drinking's sake. Now I prefer sitting around a table with friends and sipping a good beer.
In my opinion, as breweries go, Bell's is up there with some of the best. This Michigan brewery has a range of very popular craft beers as well as an interesting range of seasonal brews.
The Planet Series is their newest creation. It is based on their Batch and Wheat Series and will feature seven different beers, each inspired by a different piece of music from the composer Gustav Holst (video below). The first Planet beer will be released in August 2014 with each successive Planet debuting about every two months after, ending in July 2015.
Here's the lineup, arranged by release date and in the same order as Holst's orchestral piece:
"Mars, The Bringer of War" – Double IPA
"Venus, The Bringer of Peace" – A Blonde Ale brewed with honey, apricot, vanilla and cardamom
"Mercury, The Winged Messenger" – A Belgian Single
"Jupiter, The Bringer of Jollity" – A malt forward Brown Ale
"Saturn, The Bringer of Old Age" – A Bourbon Barrel-aged Barleywine
"Uranus, The Magician" – Black Double IPA
"Neptune, The Mystic" – A beer inspired by Dr. Bell’s Medicinal Stout, a homebrews that also helped inspire Eccentric Ale
If you aren't still mesmerized by all of the hoppy goodness, then you may be wondering "Where's Earth?" Well, if you read the names above then you probably notice that they are based on the astrological entities rather than the astronomical. And Earth gets all the beer anyway, right? So next question may be "Where's Pluto?" This isn't a Pluto-demotion issue but rather a result of the piece that The Planet Series is based on. Gustav Holst's "The Planets" predates the discovery of Pluto.
Bell's The Planet Series webpage
(story via Guysim, Geekologie, and That's Nerdalicious!)
Friday, July 4, 2014
The Bigfoot Question: A Genetic Analysis of Yeti Hair
It’s been a while since I’ve written about Bigfoot, and that’s a shame because he’s pretty fun to write about. As with many things, I like to keep it in a scientific context. That’s why I was pretty stoked to see a recent Sasquatch paper in Proceedings of the Royal Society B. A paper that takes an interesting approach: genetics.
Right off the bat the paper does not assume non-existence, both pointing out that there are numerous reports and sightings yet no bodies or recent fossils. Theories abound about what these animals are, ranging from surviving populations of collateral hominids to unlikely hybrids. As a general rule, modern science shies away from the yeti-finding field, to the point that they make believers feel rejected. Admittedly, believers have point in that science should not accept or reject anything without examining the evidence and testing hypotheses. Pretty much the definition of science, right? So that's what authors Sykes et al. do, take a scientific approach.
 The researchers collected a total of 57 Bigfoot hair samples submissions from museum and individual collections. They went about it all officially with a joint press release in May 2012 by Museum of Zoology, Lausanne and the University of Oxford. Then, to eliminate obvious non-hairs, they subjected the samples to macroscopic, microscopic and infrared fluorescence examination. Based on provenance or historic interest, thirty-seven of the samples were selected for genetic analysis. Hairs were first cleaned to remove surface contamination - just consider how many people had handled a sample, so you need to eliminate known human DNA to leave just sample DNA. The meticulously cleaned hair samples were then ground in a buffer to homogenate, incubated with proteinase K, and extracted for PCR amplification. This amplification was of the ribosomal mitrochondrial DNA 12S fragment corresponding to bps 1093-1196 of the human mitrochondrial genome, using a permissive primer combination that allows for a wide range of mammalian DNA. The results were then compared to GenBank accessions for species identification.
The researchers collected a total of 57 Bigfoot hair samples submissions from museum and individual collections. They went about it all officially with a joint press release in May 2012 by Museum of Zoology, Lausanne and the University of Oxford. Then, to eliminate obvious non-hairs, they subjected the samples to macroscopic, microscopic and infrared fluorescence examination. Based on provenance or historic interest, thirty-seven of the samples were selected for genetic analysis. Hairs were first cleaned to remove surface contamination - just consider how many people had handled a sample, so you need to eliminate known human DNA to leave just sample DNA. The meticulously cleaned hair samples were then ground in a buffer to homogenate, incubated with proteinase K, and extracted for PCR amplification. This amplification was of the ribosomal mitrochondrial DNA 12S fragment corresponding to bps 1093-1196 of the human mitrochondrial genome, using a permissive primer combination that allows for a wide range of mammalian DNA. The results were then compared to GenBank accessions for species identification. Perhaps it is important to point out what the 12S mitochondrial DNA is and how it works. Even within fur-bearing species, there is a large amount of variation in hair appearance that can be identified under the microscope to determine species. But, in the absence of an experienced hair examiner (yes, those exist), a reliable, alternative analysis must be used. This analysis comes in the form of highly conserved mitochondrial DNA regions, these are particular sequences that have been maintained by evolution despite speciation, probably because they are functional. Mitochondrial 12S ribosomal RNA has an amplification size that renders it useful for even problematic and/or degraded samples. Highly conserved primer regions and the high nucleotide species diversity present within the portion of the 12S gene examined allows for identification at least to genus and often species. Studies examining the extent of 12S homology within and between species have shown a high degree of confidence in the test's ability to match species from biological samples, usually hair. This includes primate homologies like the chimpanzee, who shares a 98% homology with the human 12S region, Gorilla (97%) and rhesus macaque (90%). These studies have shown that it is unlikely that a non-human primate hair could be confused with human hair using this system.
Perhaps it is important to point out what the 12S mitochondrial DNA is and how it works. Even within fur-bearing species, there is a large amount of variation in hair appearance that can be identified under the microscope to determine species. But, in the absence of an experienced hair examiner (yes, those exist), a reliable, alternative analysis must be used. This analysis comes in the form of highly conserved mitochondrial DNA regions, these are particular sequences that have been maintained by evolution despite speciation, probably because they are functional. Mitochondrial 12S ribosomal RNA has an amplification size that renders it useful for even problematic and/or degraded samples. Highly conserved primer regions and the high nucleotide species diversity present within the portion of the 12S gene examined allows for identification at least to genus and often species. Studies examining the extent of 12S homology within and between species have shown a high degree of confidence in the test's ability to match species from biological samples, usually hair. This includes primate homologies like the chimpanzee, who shares a 98% homology with the human 12S region, Gorilla (97%) and rhesus macaque (90%). These studies have shown that it is unlikely that a non-human primate hair could be confused with human hair using this system.Now knowing all of this, back to the results of the Bigfoot study. Despite multiple attempts, seven of the samples yielded no DNA sequences, leaving the researchers with 30 samples. These 30 samples were each matched to a known species. Ten belonged to various bear species, four were cows, four were horse, four were wolves/dogs, two were raccoons, one was a deer, one a Malaysian tapir, one a sheep, one a serow, and one was human (exact match).
There has been quite a few articles in the news about this study, and that’s good because this paper is a nice example of using hard science to test a theory. It is also works towards bridging the gap between two rather disparate groups of people. So kudos to you Sykes et al.
A nice write-up from Science News "'Bigfoot' samples analyzed in lab"
For more on 12S see an article in Forensic Magazine titled "Easy Species DNA Identification for the Forensic Laboratory Using 12S Mitochondrial DNA"
(images via WhoFortedBlog, NewEngland BioLabs, Nature Reviews Genetic paper DOI:10.1038/nrg1606, respectively)
Subscribe to:
Posts (Atom)





